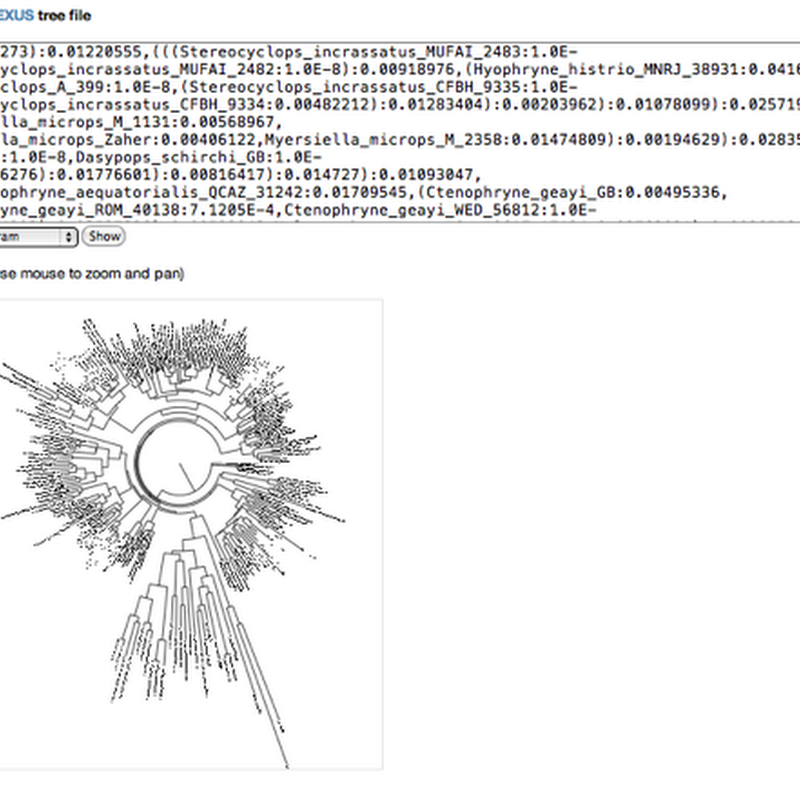
Say what you will about Elsevier, they are certainly exploring ways to re-imagine the scientific article. In a comment on an earlier post Fabian Schreiber pointed out that Elsevier have released an app to display phylogenies in articles they publish. The app is based on jsPhyloSVGand is described here.