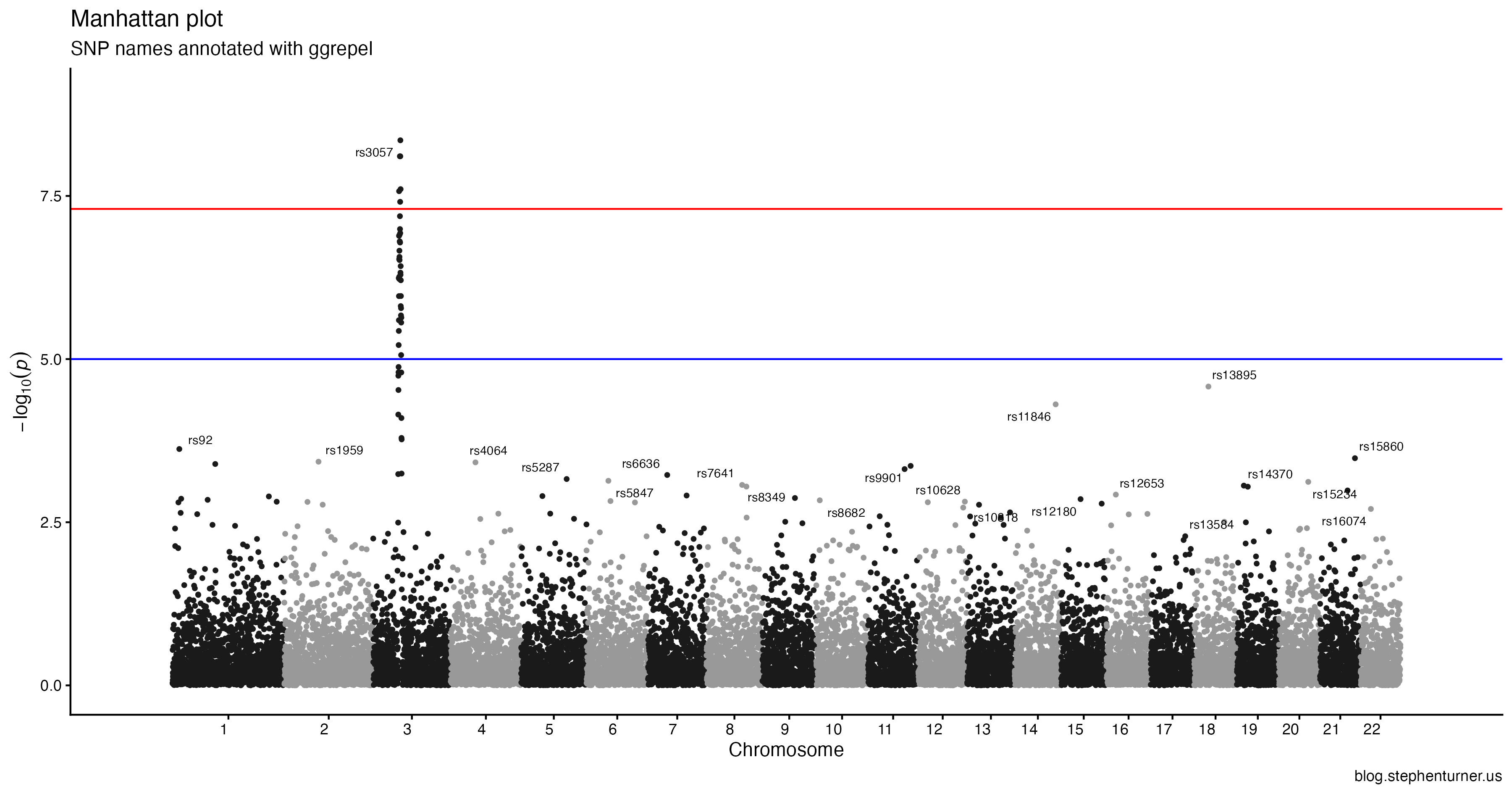
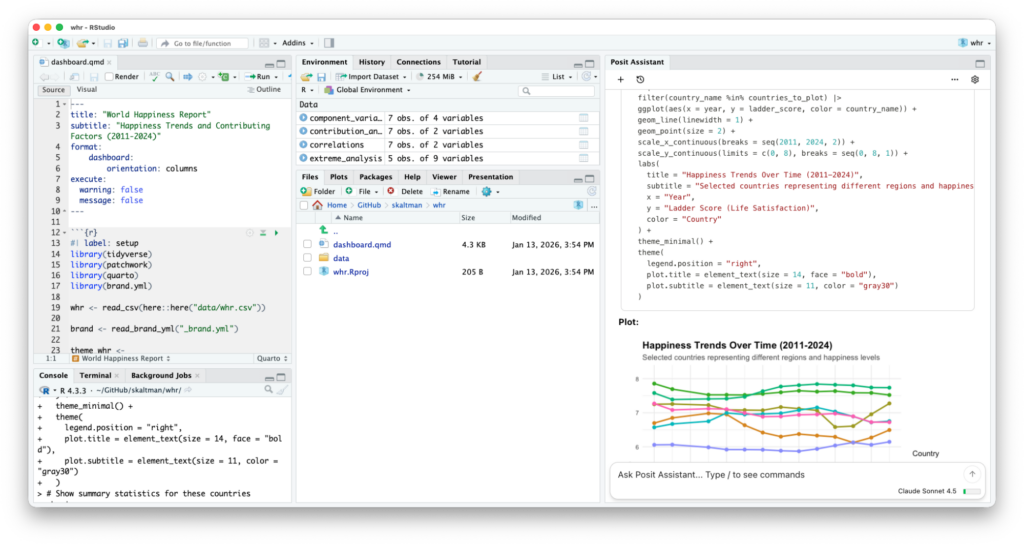
Vibe-updating my old qqman R package to ggplot2 with plan+execute.

Vibe-updating my old qqman R package to ggplot2 with plan+execute.
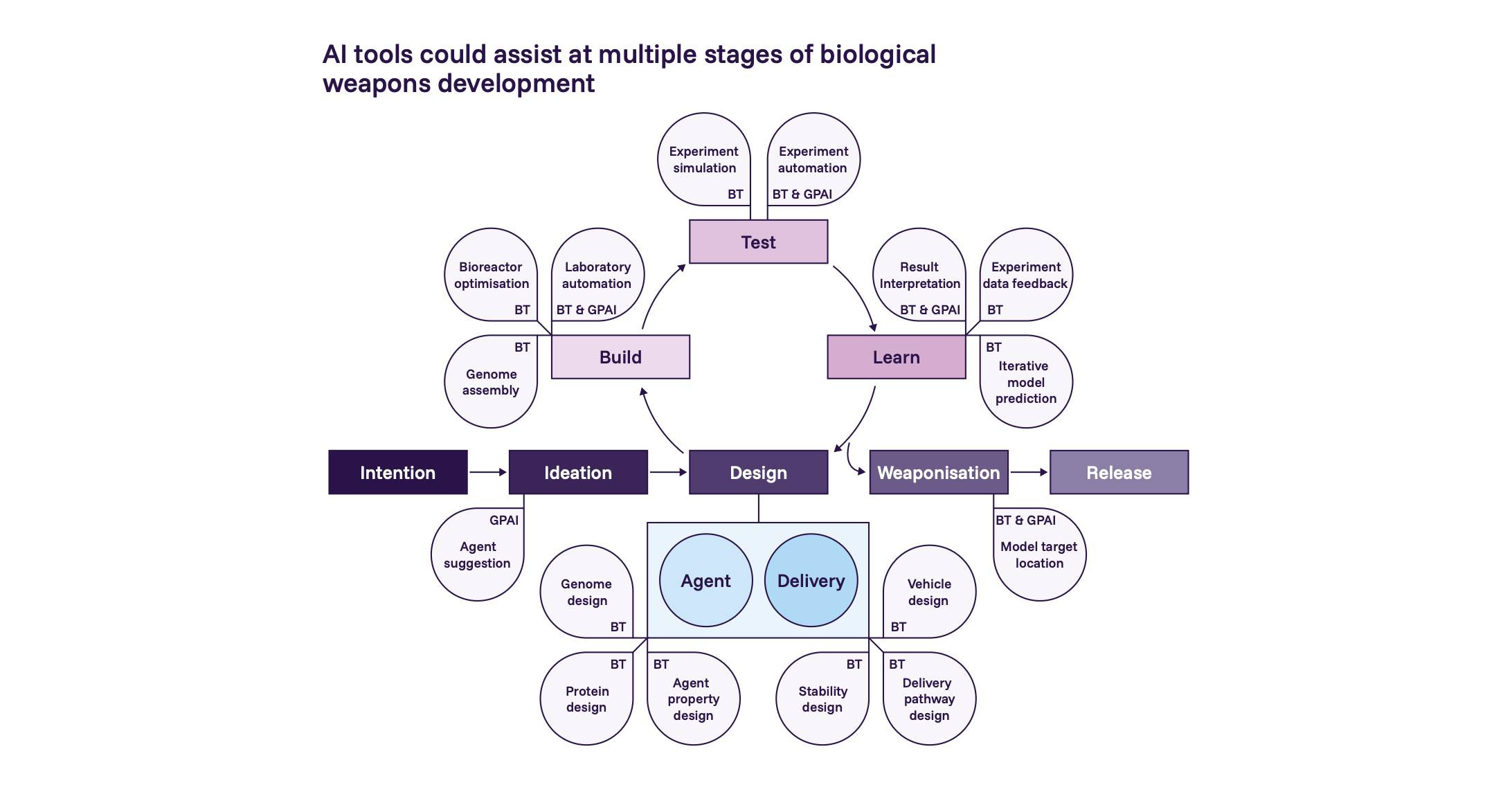
Notes from the 2026 International AI Safety Report

What's "tacit knowledge" in biosecurity, and is it really a barrier? A look at the 2025 RAND paper "Contemporary Foundation AI Models Increase Biological Weapons Risk." 1.7k words, 8 min reading time.
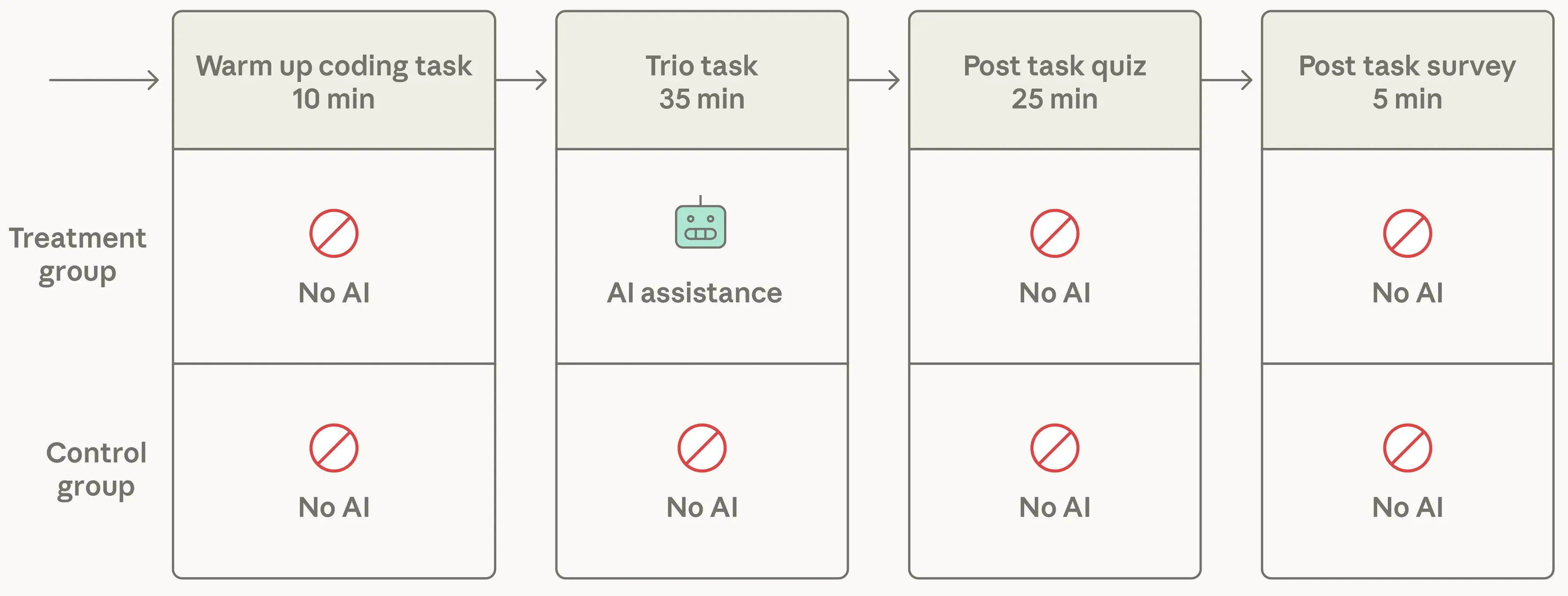
Research from scientists at Anthropic suggests productivity gains come at the cost of skill development.
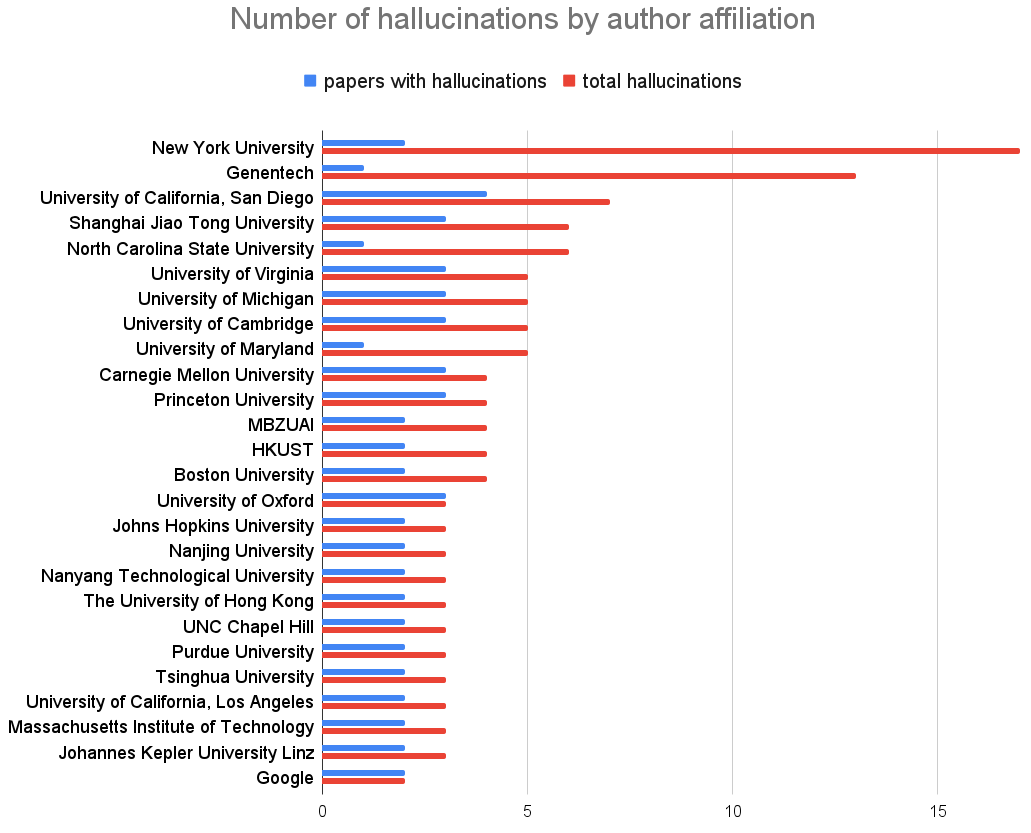
Global Biodiversity Framework, Colossal+Conservation, DARPA GUARDIAN, NTI on AIxBio, NeurIPS hallucitations, Claude in Excel, AI@UVA, Amodei essay, R updates (R Weekly, R Data Scientist), papers.
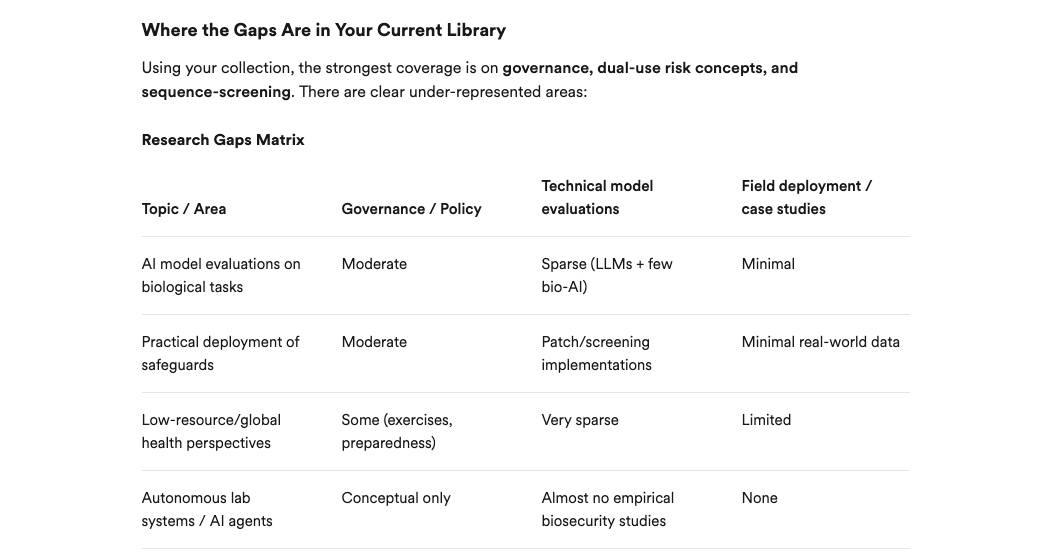
Sync your Zotero library to Consensus AI to ask questions about papers you've saved, and find and fill in gaps in your collections.

Whether this is your first conference talk or your fiftieth, we’re looking for speakers from a variety of backgrounds and experience levels
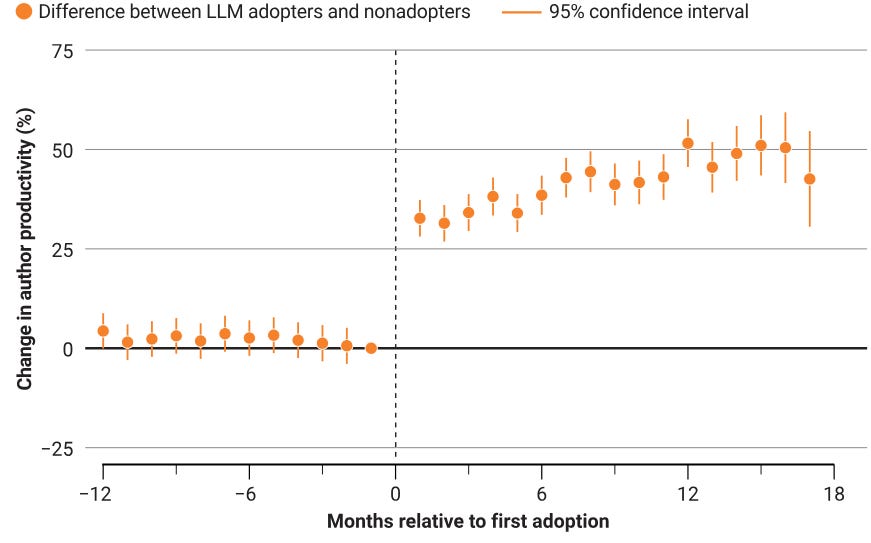
With the production process rapidly evolving, science policy must consider how institutions could evolve

If you read Paired Ends because you care about how biology, technology, and society co-evolve, this story was written for you.
AIxBio, Claude's Constitution, UK AI+Science, exams with chatbots, R updates (Posit, R Data Scientist, R Weekly), reading what Adam Kucharski is reading, AI vs reality at Salesforce, papers+preprints.
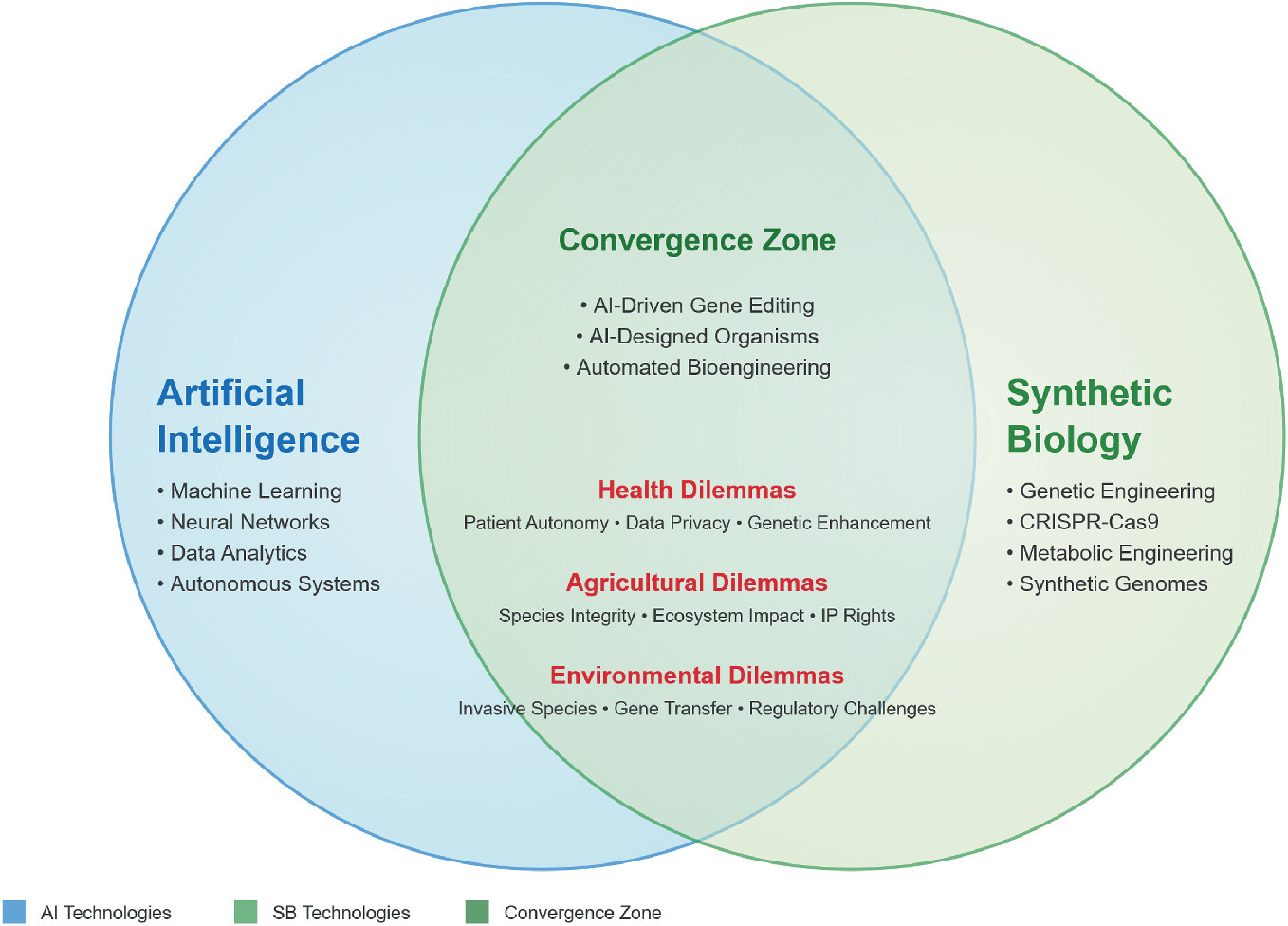
Part 2 summarizing chapters from "Biotechnology and AI: Technological Convergence and Information Hazards" from the NATO Advanced Research Workshop on AI and biotechnology convergence